Travels in DC: Yuanhang Liu at the BIBM Conference
Yuanhang Liu (Genetics, Genomics & Development Track) was selected to receive a GSBS travel award based on his
“excellent academic record and standing in the scientific community.” The article below is a summary of his time at the BIBM conference.
Firstly, thanks again to GSBS travel award review committee for offering me travel award to attend the IEEE International Conference on Bioinformatics and Biomedicine (BIBM) conference in D.C. I consider myself really lucky to be in D.C and met lot of interesting people here.
The BIBM conference provides a leading forum for disseminating the latest research in bioinformatics and health informatics. It brings together academic and industrial scientists from computer science, biology, chemistry,
medicine, mathematics, and statistics. BIBM was held in Washington D.C this year, which is very close to the National Institute of Health (NIH) located in Maryland.

When I was at BIBM, I found many of the talks to be interesting. The first was a talk by Associate Director for Data Science (ADDS) at the National
Institutes of Health Dr. Philip E. Bourne, talked about his perceptive of the future of biomedical sciences and NIH’s recent effort in advancing our understanding of human health and disease.
He pointed out that lack of appropriate tools, poor data accessibility, and insufficient training, are major impediments to rapid translational impact.
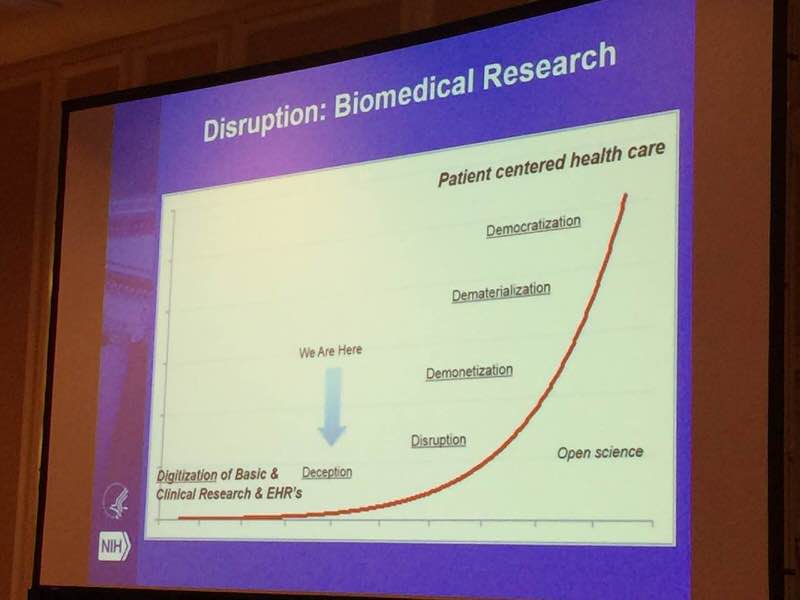
To meet this challenge, NIH launched the Big Data to Knowledge (BD2K) initiative in 2012.
Currently, NIH has already established 13 centers for big data computing and data coordination and integration, including the broad institute and The University of California Santa Cruz (UCSC), etc. Dr. Bourne also talked about how this community can further engage in these activities.
Another talk I found interesting was Dr. Wei Wang’s talk regarding RNA-seq analysis.Dr. Wei Wang is professor in the Department of Computer Science at University of California at Los Angeles and the director of the Scalable Analytics Institute (ScAi).
She talked about their recent work regarding a novel RNA-Seq quantification method, RNA-Skim, which partitions the transcriptome into disjoint transcript clusters based on sequence similarity, and introduces the notion of sig-mers, which are a special type of k-mers uniquely associated with each cluster. This enables RNA-Skim to perform transcript quantification on each cluster independently, reducing a complex optimization problem into smaller optimization tasks that can be run in parallel.

As a result, RNA-Skim is able to finish transcriptome quantification in under
10 min per sample by using just a single thread on a commodity computer, which
represents over 100 speedup over the state-of-the-art alignment-based methods,
while delivering comparable or higher accuracy
During my stay, I was lucky to meet one of the attendees that came from the same university as me in China. Currently, he is a research scientist at the Georgia Institute of Technology. We shared opinions about the future of biomedical science and what we can do about it. It was really a great
experience!
I was also able to visit the several museums and memorials during my stay. I took some pictures during the night, which, seemed to be more attractive to me. Enjoy!
The “Beyond The Bench” series features articles written by students and postdoctoral fellows at the Graduate School of Biomedical Sciences at The University of Texas Health Science Center San Antonio.



